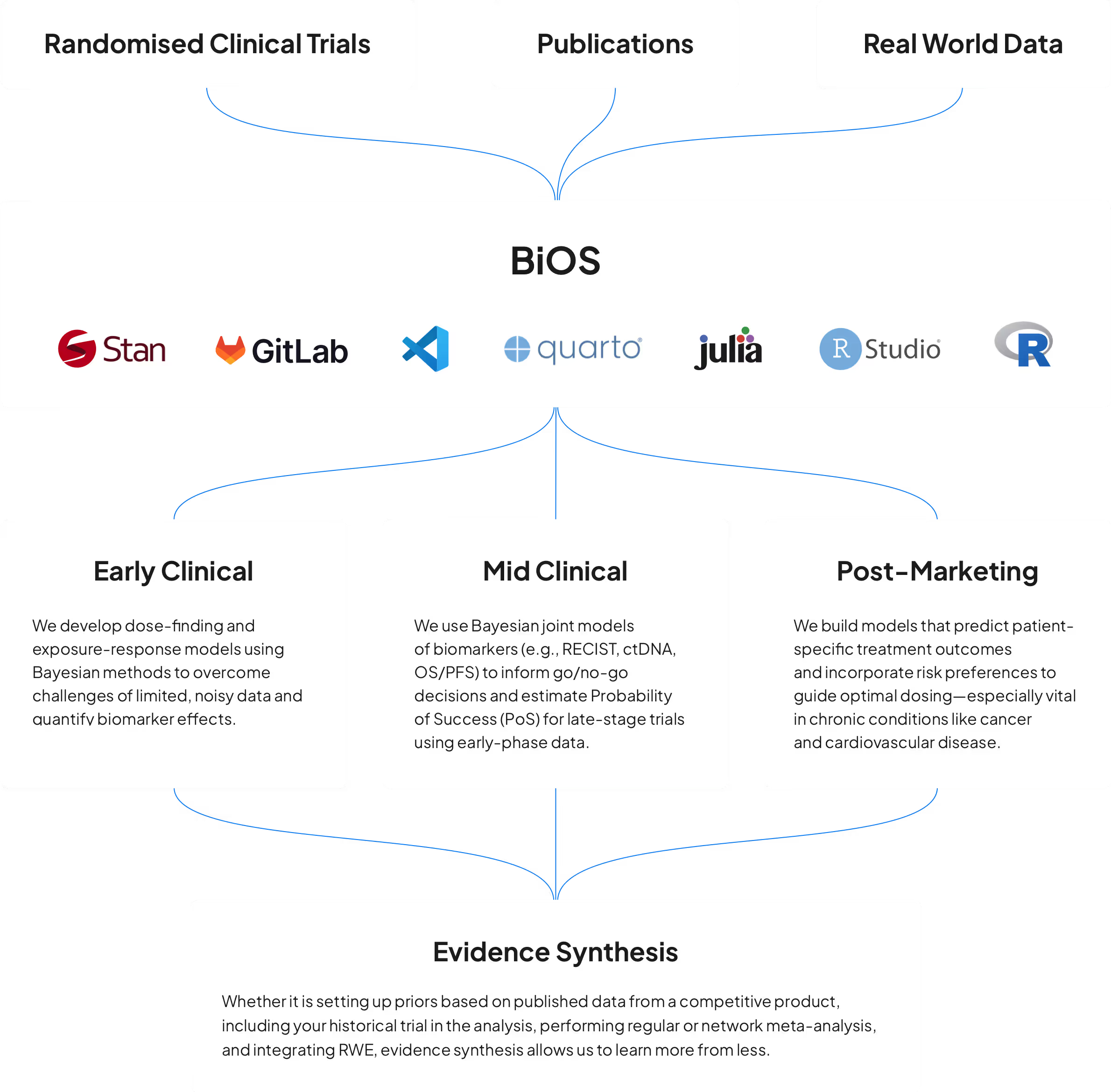
Big data is all the rage, but what do you do when you only have a few patients and few observations? Our models work well in the small data regime, such as early clinical trials and rare diseases, where we need to take advantage of prior knowledge from previous studies, scientific publications, and other external data sources.